The Clustered Variance module adjusts standard errors for clustering. For example, replicating a dataset 100 times should not increase the precision of parameter estimates, but performing this procedure with the IID assumption will actually do this. Another example is in economics of education research, it is reasonable to expect that the error terms for children in the same class are not independent. Clustering standard errors can correct for this.
The MADlib Clustered Variance module includes functions to calculate linear, logistic, and multinomial logistic regression problems.
The clustered variance linear regression training function has the following syntax.
clustered_variance_linregr ( source_table,
out_table,
dependent_varname,
independent_varname,
clustervar,
grouping_cols
)
Arguments
TEXT. The name of the table containing the input data.
VARCHAR. Name of the generated table containing the output model. The output table contains the following columns.
| coef | DOUBLE PRECISION[]. Vector of the coefficients of the regression. |
|---|---|
| std_err | DOUBLE PRECISION[]. Vector of the standard error of the coefficients. |
| t_stats | DOUBLE PRECISION[]. Vector of the t-stats of the coefficients. |
| p_values | DOUBLE PRECISION[]. Vector of the p-values of the coefficients. |
A summary table named <out_table>_summary is also created, which is the same as the summary table created by linregr_train function. Please refer to the documentation for linear regression for details.
The clustered variance logistic regression training function has the following syntax.
clustered_variance_logregr( source_table,
out_table,
dependent_varname,
independent_varname,
clustervar,
grouping_cols,
max_iter,
optimizer,
tolerance,
verbose_mode
)
Arguments
VARCHAR. Name of the generated table containing the output model. The output table has the following columns:
| coef | Vector of the coefficients of the regression. |
|---|---|
| std_err | Vector of the standard error of the coefficients. |
| z_stats | Vector of the z-stats of the coefficients. |
| p_values | Vector of the p-values of the coefficients. |
A summary table named <out_table>_summary is also created, which is the same as the summary table created by logregr_train function. Please refer to the documentation for logistic regression for details.
clustered_variance_mlogregr( source_table,
out_table,
dependent_varname,
independent_varname,
cluster_varname,
ref_category,
grouping_cols,
optimizer_params,
verbose_mode
)
Arguments
TEXT. The name of the table where the regression model will be stored. The output table has the following columns:
| category | The category. |
|---|---|
| ref_category | The refererence category used for modeling. |
| coef | Vector of the coefficients of the regression. |
| std_err | Vector of the standard error of the coefficients. |
| z_stats | Vector of the z-stats of the coefficients. |
| p_values | Vector of the p-values of the coefficients. |
A summary table named <out_table>_summary is also created, which is the same as the summary table created by mlogregr_train function. Please refer to the documentation for multinomial logistic regression for details.
The clustered robust variance estimator function for the Cox Proportional Hazards model has the following syntax.
clustered_variance_coxph(model_table, output_table, clustervar)
Arguments
| coef | FLOAT8[]. Vector of the coefficients. |
|---|---|
| loglikelihood | FLOAT8. Log-likelihood value of the MLE estimate. |
| std_err | FLOAT8[]. Vector of the standard error of the coefficients. |
| clustervar | TEXT. A comma-separated list of columns to use as cluster variables. |
| clustered_se | FLOAT8[]. Vector of the robust standard errors of the coefficients. |
| clustered_z | FLOAT8[]. Vector of the robust z-stats of the coefficients. |
| clustered_p | FLOAT8[]. Vector of the robust p-values of the coefficients. |
| hessian | FLOAT8[]. The Hessian matrix. |
SELECT madlib.clustered_variance_linregr();
DROP TABLE IF EXISTS out_table;
SELECT madlib.clustered_variance_linregr( 'abalone',
'out_table',
'rings',
'ARRAY[1, diameter, length, width]',
'sex',
NULL
);
SELECT * FROM out_table;
SELECT madlib.clustered_variance_logregr();
DROP TABLE IF EXISTS out_table;
SELECT madlib.clustered_variance_logregr( 'abalone',
'out_table',
'rings < 10',
'ARRAY[1, diameter, length, width]',
'sex'
);
SELECT * FROM out_table;
SELECT madlib.clustered_variance_mlogregr();
DROP TABLE IF EXISTS out_table;
SELECT madlib.clustered_variance_mlogregr( 'abalone',
'out_table',
'CASE WHEN rings < 10 THEN 1 ELSE 0 END',
'ARRAY[1, diameter, length, width]',
'sex',
0
);
SELECT * FROM out_table;
DROP TABLE IF EXISTS lung_cl_out;
DROP TABLE IF EXISTS lung_out;
DROP TABLE IF EXISTS lung_out_summary;
SELECT madlib.coxph_train('lung',
'lung_out',
'time',
'array[age, "ph.ecog"]',
'TRUE',
NULL,
NULL);
SELECT madlib.clustered_variance_coxph('lung_out',
'lung_cl_out',
'"ph.karno"');
SELECT * FROM lung_cl_out;
Assume that the data can be separated into 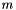 clusters. Usually this can be done by grouping the data table according to one or multiple columns.
clusters. Usually this can be done by grouping the data table according to one or multiple columns.
The estimator has a similar form to the usual sandwich estimator
![\[ S(\vec{c}) = B(\vec{c}) M(\vec{c}) B(\vec{c}) \]](form_315.png)
The bread part is the same as Huber-White sandwich estimator
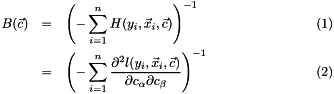
where 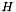 is the hessian matrix, which is the second derivative of the target function
is the hessian matrix, which is the second derivative of the target function
![\[ L(\vec{c}) = \sum_{i=1}^n l(y_i, \vec{x}_i, \vec{c})\ . \]](form_318.png)
The meat part is different
![\[ M(\vec{c}) = \bf{A}^T\bf{A} \]](form_319.png)
where the 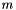 -th row of
-th row of 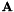 is
is
![\[ A_m = \sum_{i\in G_m}\frac{\partial l(y_i,\vec{x}_i,\vec{c})}{\partial \vec{c}} \]](form_321.png)
where 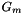 is the set of rows that belong to the same cluster.
is the set of rows that belong to the same cluster.
We can compute the quantities of 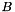 and
and 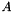 for each cluster during one scan through the data table in an aggregate function. Then sum over all clusters to the full
for each cluster during one scan through the data table in an aggregate function. Then sum over all clusters to the full 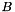 and
and 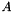 in the outside of the aggregate function. At last, the matrix mulplitications are done in a separate function on the master node.
in the outside of the aggregate function. At last, the matrix mulplitications are done in a separate function on the master node.
When multinomial logistic regression is computed before the multinomial clustered variance calculation, it uses a default reference category of zero and the regression coefficients are included in the output table. The regression coefficients in the output are in the same order as multinomial logistic regression function, which is described below. For a problem with 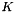 dependent variables
dependent variables 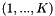 and
and 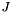 categories
categories 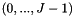 , let
, let 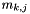 denote the coefficient for dependent variable
denote the coefficient for dependent variable 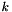 and category
and category 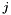 . The output is
. The output is 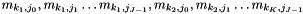 . The order is NOT CONSISTENT with the multinomial regression marginal effect calculation with function marginal_mlogregr. This is deliberate because the interfaces of all multinomial regressions (robust, clustered, ...) will be moved to match that used in marginal.
. The order is NOT CONSISTENT with the multinomial regression marginal effect calculation with function marginal_mlogregr. This is deliberate because the interfaces of all multinomial regressions (robust, clustered, ...) will be moved to match that used in marginal.
[1] Standard, Robust, and Clustered Standard Errors Computed in R, http://diffuseprior.wordpress.com/2012/06/15/standard-robust-and-clustered-standard-errors-computed-in-r/
File clustered_variance_coxph.sql_in documenting the clustered variance for Cox proportional hazards SQL functions.