This module implements elastic net regularization [1] for linear and logistic regression. Regularization is a technique often used to prevent overfitting.
elastic_net_train( tbl_source,
tbl_result,
col_dep_var,
col_ind_var,
regress_family,
alpha,
lambda_value,
standardize,
grouping_col,
optimizer,
optimizer_params,
excluded,
max_iter,
tolerance
)
Arguments
TEXT. The name of the table containing the training data.
TEXT. Name of the output table containing output model. The output table produced by the elastic_net_train() function has the following columns:
| regress_family | The regression type: 'gaussian' or 'binomial'. |
|---|---|
| features | Array of features (independent variables) passed to the algorithm. |
| features_selected | Array of features selected by the algorithm. |
| coef_nonzero | Coefficients of the selected features. |
| coef_all | Coefficients of all features, both selected and unselected. |
| intercept | Intercept for the model. |
| log_likelihood | Log of the likelihood value produced by the algorithm. |
| standardize | BOOLEAN. If data has been normalized, will be set to TRUE. |
| iteration_run | The number of iterations executed. |
TEXT. An expression for the dependent variable.
col_dep_var = 'log(y+1)', and col_ind_var = 'array[exp(x[1]), x[2], 1/(1+x[3])]'. In the binomial case, you can use a Boolean expression, for example, col_dep_var = 'y < 0'.TEXT. An expression for the independent variables. Use '*' to specify all columns of tbl_source except those listed in the excluded string described below. If col_dep_var is a column name, it is automatically excluded from the independent variables. However, if col_dep_var is a valid PostgreSQL expression, any column names used within the expression are only excluded if they are explicitly listed in the excluded argument. Therefore, it is a good idea to add all column names involved in the dependent variable expression to the excluded string.
TEXT. For regression type, specify either 'gaussian' ('linear') or 'binomial' ('logistic').
FLOAT8. Elastic net control parameter with a value in the range [0, 1]. A value of 1 means L1 regularization, and a value of 0 means L2 regularization.
FLOAT8. Regularization parameter (must be positive).
BOOLEAN, default: TRUE. Whether to normalize the data or not. Setting to TRUE usually yields better results and faster convergence.
TEXT, default: NULL. A single column or a list of comma-separated columns that divides the input data into discrete groups, resulting in one regression per group. When this value is NULL, no grouping is used and a single model is generated for all data.
TEXT, default: 'fista'. Name of optimizer, either 'fista' or 'igd'. FISTA [2] is an algorithm with a fast global rate of convergence for solving linear inverse problems. Incremental gradient descent (IGD) is a stochastic approach to minimizing an objective function [4].
TEXT, default: NULL. Optimizer parameters, delimited with commas. These parameters differ depending on the value of optimizer parameter. See the descriptions below for details.
TEXT, default: NULL. If the col_ind_var input is '*' then excluded can be provided as a comma-delimited list of column names that are to be excluded from the features. For example, 'col1, col2'. If the col_ind_var is an array, excluded must be a list of the integer array positions to exclude, for example '1,2'. If this argument is NULL or an empty string, no columns are excluded.
INTEGER, default: 1000. The maximum number of iterations allowed.
For optimizer_params, there are several parameters that can be supplied in a string containing a comma-delimited list of name-value pairs . All of these named parameters are optional and use the format "<param_name> = <value>".
The parameters described below are organized by category: warmup, cross validation and optimization.
Warmup parameters
$$
warmup = <value>,
warmup_lambdas = <value>,
warmup_lambda_no = <value>,
warmup_tolerance = <value>
$$
Default: FALSE. If warmup is TRUE, a series of strictly descending lambda values are used, which end with the lambda value that the user wants to calculate. A larger lambda gives a sparser solution, and the sparse solution is then used as the initial guess for the next lambda's solution, which can speed up the computation for the next lambda. For larger data sets, this can sometimes accelerate the whole computation and may in fact be faster than computation with only a single lambda value.
Default: NULL. Set of lambda values to use when warmup is TRUE. The default is NULL, which means that lambda values will be automatically generated.
Default: 15. Number of lambda values used in warm-up. If warmup_lambdas is not NULL, this value is overridden by the number of provided lambda values.
Cross validation parameters
$$
n_folds = <value>,
validation_result = <value>,
lambda_value = <value>,
n_lambdas = <value>,
alpha = <value>
$$
Hyperparameter optimization can be carried out using the built-in cross validation mechanism, which is activated by assigning a value greater than 1 to the parameter n_folds. Misclassification error is used for classification and mean squared error is used for regression.
The values of a parameter to cross validate should be provided in a list. For example, to regularize with the L1 norm and use a lambda value from the set {0.3, 0.4, 0.5}, include 'lambda_value={0.3, 0.4, 0.5}'. Note that the use of '{}' and '[]' are both valid here.
Default: 0. Number of folds (k). Must be at least 2 to activate cross validation. If a value of k > 2 is specified, each fold is then used as a validation set once, while the other k - 1 folds form the training set.
Default: NULL. Name of the table to store the cross validation results, including the values of parameters and their averaged error values. The table is only created if the name is not NULL.
Default: NULL. Set of regularization values to be used for cross validation. The default is NULL, which means that lambda values will be automatically generated.
Default: 15. Number of lambdas to cross validate over. If a list of lambda values is not provided in the lambda_value set above, this parameter can be used to autogenerate the set of lambdas. If the lambda_value set is not NULL, this value is overridden by the number of provided lambda values.
Optimizer parameters
FISTA Parameters
$$
max_stepsize = <value>,
eta = <value>,
use_active_set = <value>,
activeset_tolerance = <value>,
random_stepsize = <value>
$$
Default: 4.0. Initial backtracking step size. At each iteration, the algorithm first tries stepsize = max_stepsize, and if it does not work out, it then tries a smaller step size, stepsize = stepsize/eta, where eta must be larger than 1. At first glance, this seems to perform repeated iterations for even one step, but using a larger step size actually greatly increases the computation speed and minimizes the total number of iterations. A careful choice of max_stepsize can decrease the computation time by more than 10 times.
Default: 2.0 If stepsize does not work, stepsize/eta is tried. Must be greater than 1.
Default: FALSE. If use_active_set is TRUE, an active-set method is used to speed up the computation. Considerable speedup is obtained by organizing the iterations around the active set of features—those with nonzero coefficients. After a complete cycle through all the variables, we iterate only on the active set until convergence. If another complete cycle does not change the active set, we are done. Otherwise, the process is repeated.
The value of tolerance used during active set calculation. The default value is the same as the tolerance argument described above.
IGD parameters
$$
stepsize = <value>,
step_decay = <value>,
threshold = <value>,
parallel = <value>
$$
The default is 0.01.
The actual stepsize used for current step is (previous stepsize) / exp(step_decay). The default value is 0, which means that a constant stepsize is used in IGD.
Default: 1e-10. When a coefficient is really small, set this coefficient to be 0.
Due to the stochastic nature of SGD, we can only obtain very small values for the fitting coefficients. Therefore, threshold is needed at the end of the computation to screen out tiny values and hard-set them to zeros. This is accomplished as follows: (1) multiply each coefficient with the standard deviation of the corresponding feature; (2) compute the average of absolute values of re-scaled coefficients; (3) divide each rescaled coefficient with the average, and if the resulting absolute value is smaller than threshold, set the original coefficient to zero.
Whether to run the computation on multiple segments. The default is TRUE.
SGD is a sequential algorithm in nature. When running in a distributed manner, each segment of the data runs its own SGD model and then the models are averaged to get a model for each iteration. This averaging might slow down the convergence speed, but it affords the ability to process large datasets on a cluster of machines. This algorithm, therefore, provides the parallel option to allow you to choose whether to do parallel computation.
The prediction function returns a double value for the Gaussian family and a Boolean value for the Binomial family.
The predict function has the following syntax (elastic_net_gaussian_predict() and elastic_net_binomial_predict()):
elastic_net_<family>_predict(
coefficients,
intercept,
ind_var
)
Arguments
For the binomial family, there is a function (elastic_net_binomial_prob()) that outputs the probability of the instance being TRUE:
elastic_net_binomial_prob(
coefficients,
intercept,
ind_var
)
Alternatively, you can use another prediction function that stores the prediction result in a table (elastic_net_predict()). This is useful if you want to use elastic net together with the general cross validation function.
elastic_net_predict( tbl_model,
tbl_new_sourcedata,
col_id,
tbl_predict
)
Arguments
You do not need to specify whether the model is "linear" or "logistic" because this information is already included in the tbl_model table.
SELECT madlib.elastic_net_train();
DROP TABLE IF EXISTS houses;
CREATE TABLE houses ( id INT,
tax INT,
bedroom INT,
bath FLOAT,
price INT,
size INT,
lot INT,
zipcode INT);
INSERT INTO houses (id, tax, bedroom, bath, price, size, lot, zipcode) VALUES
(1 , 590 , 2 , 1 , 50000 , 770 , 22100 , 94301),
(2 , 1050 , 3 , 2 , 85000 , 1410 , 12000 , 94301),
(3 , 20 , 3 , 1 , 22500 , 1060 , 3500 , 94301),
(4 , 870 , 2 , 2 , 90000 , 1300 , 17500 , 94301),
(5 , 1320 , 3 , 2 , 133000 , 1500 , 30000 , 94301),
(6 , 1350 , 2 , 1 , 90500 , 820 , 25700 , 94301),
(7 , 2790 , 3 , 2.5 , 260000 , 2130 , 25000 , 94301),
(8 , 680 , 2 , 1 , 142500 , 1170 , 22000 , 94301),
(9 , 1840 , 3 , 2 , 160000 , 1500 , 19000 , 94301),
(10 , 3680 , 4 , 2 , 240000 , 2790 , 20000 , 94301),
(11 , 1660 , 3 , 1 , 87000 , 1030 , 17500 , 94301),
(12 , 1620 , 3 , 2 , 118600 , 1250 , 20000 , 94301),
(13 , 3100 , 3 , 2 , 140000 , 1760 , 38000 , 94301),
(14 , 2070 , 2 , 3 , 148000 , 1550 , 14000 , 94301),
(15 , 650 , 3 , 1.5 , 65000 , 1450 , 12000 , 94301),
(16 , 770 , 2 , 2 , 91000 , 1300 , 17500 , 76010),
(17 , 1220 , 3 , 2 , 132300 , 1500 , 30000 , 76010),
(18 , 1150 , 2 , 1 , 91100 , 820 , 25700 , 76010),
(19 , 2690 , 3 , 2.5 , 260011 , 2130 , 25000 , 76010),
(20 , 780 , 2 , 1 , 141800 , 1170 , 22000 , 76010),
(21 , 1910 , 3 , 2 , 160900 , 1500 , 19000 , 76010),
(22 , 3600 , 4 , 2 , 239000 , 2790 , 20000 , 76010),
(23 , 1600 , 3 , 1 , 81010 , 1030 , 17500 , 76010),
(24 , 1590 , 3 , 2 , 117910 , 1250 , 20000 , 76010),
(25 , 3200 , 3 , 2 , 141100 , 1760 , 38000 , 76010),
(26 , 2270 , 2 , 3 , 148011 , 1550 , 14000 , 76010),
(27 , 750 , 3 , 1.5 , 66000 , 1450 , 12000 , 76010);
DROP TABLE IF EXISTS houses_en, houses_en_summary;
SELECT madlib.elastic_net_train( 'houses', -- Source table
'houses_en', -- Result table
'price', -- Dependent variable
'array[tax, bath, size]', -- Independent variable
'gaussian', -- Regression family
0.5, -- Alpha value
0.1, -- Lambda value
TRUE, -- Standardize
NULL, -- Grouping column(s)
'fista', -- Optimizer
'', -- Optimizer parameters
NULL, -- Excluded columns
10000, -- Maximum iterations
1e-6 -- Tolerance value
);
-- Turn on expanded display to make it easier to read results. \x on SELECT * FROM houses_en;Result:
-[ RECORD 1 ]-----+-------------------------------------------
family | gaussian
features | {tax,bath,size}
features_selected | {tax,bath,size}
coef_nonzero | {22.785201806,10707.9664343,54.7959774173}
coef_all | {22.785201806,10707.9664343,54.7959774173}
intercept | -7798.71393905
log_likelihood | -512248641.971
standardize | t
iteration_run | 10000
\x off
SELECT id, price, predict, price - predict AS residual
FROM (
SELECT
houses.*,
madlib.elastic_net_gaussian_predict(
m.coef_all, -- Coefficients
m.intercept, -- Intercept
ARRAY[tax,bath,size] -- Features (corresponding to coefficients)
) AS predict
FROM houses, houses_en m) s
ORDER BY id;
Result: id | price | predict | residual ----+--------+------------------+------------------- 1 | 50000 | 58545.391894031 | -8545.391894031 2 | 85000 | 114804.077663003 | -29804.077663003 3 | 22500 | 61448.835664388 | -38948.835664388 4 | 90000 | 104675.17768007 | -14675.17768007 5 | 133000 | 125887.70644358 | 7112.29355642 6 | 90500 | 78601.843595366 | 11898.156404634 7 | 260000 | 199257.358231079 | 60742.641768921 8 | 142500 | 82514.559377081 | 59985.440622919 9 | 160000 | 137735.93215082 | 22264.06784918 10 | 240000 | 250347.627648647 | -10347.627648647 11 | 87000 | 97172.428263539 | -10172.428263539 12 | 118600 | 119024.150628605 | -424.150628604999 13 | 140000 | 180692.127913358 | -40692.127913358 14 | 148000 | 156424.249824545 | -8424.249824545 15 | 65000 | 102527.938104575 | -37527.938104575 16 | 91000 | 102396.67273637 | -11396.67273637 17 | 132300 | 123609.20149988 | 8690.79850012 18 | 91100 | 74044.833707966 | 17055.166292034 19 | 260011 | 196978.853287379 | 63032.146712621 20 | 141800 | 84793.064320781 | 57006.935679219 21 | 160900 | 139330.88561141 | 21569.11438859 22 | 239000 | 248524.823693687 | -9524.82369368701 23 | 81010 | 95805.325297319 | -14795.325297319 24 | 117910 | 118340.599145495 | -430.599145494998 25 | 141100 | 182970.632857058 | -41870.632857058 26 | 148011 | 160981.259711945 | -12970.259711945 27 | 66000 | 104806.443048275 | -38806.443048275
DROP TABLE IF EXISTS houses_en1, houses_en1_summary;
SELECT madlib.elastic_net_train( 'houses', -- Source table
'houses_en1', -- Result table
'price', -- Dependent variable
'array[tax, bath, size]', -- Independent variable
'gaussian', -- Regression family
0.5, -- Alpha value
0.1, -- Lambda value
TRUE, -- Standardize
'zipcode', -- Grouping column(s)
'fista', -- Optimizer
'', -- Optimizer parameters
NULL, -- Excluded columns
10000, -- Maximum iterations
1e-6 -- Tolerance value
);
-- Turn on expanded display to make it easier to read results. \x on SELECT * FROM houses_en1;Result:
-[ RECORD 1 ]-----+--------------------------------------------
zipcode | 94301
family | gaussian
features | {tax,bath,size}
features_selected | {tax,bath,size}
coef_nonzero | {27.0542096962,12351.5244083,47.5833289771}
coef_all | {27.0542096962,12351.5244083,47.5833289771}
intercept | -7191.19791597
log_likelihood | -519199964.967
standardize | t
iteration_run | 10000
-[ RECORD 2 ]-----+--------------------------------------------
zipcode | 76010
family | gaussian
features | {tax,bath,size}
features_selected | {tax,bath,size}
coef_nonzero | {15.6325953499,10166.6608469,57.8689916035}
coef_all | {15.6325953499,10166.6608469,57.8689916035}
intercept | 513.912201627
log_likelihood | -538806528.45
standardize | t
iteration_run | 10000
\x off
SELECT madlib.elastic_net_predict(
'houses_en1', -- Model table
'houses', -- New source data table
'id', -- Unique ID associated with each row
'houses_en1_prediction' -- Table to store prediction result
);
SELECT houses.id,
houses.price,
houses_en1_prediction.prediction,
houses.price - houses_en1_prediction.prediction AS residual
FROM houses_en1_prediction, houses
WHERE houses.id = houses_en1_prediction.id ORDER BY id;
DROP TABLE IF EXISTS houses_en2, houses_en2_summary;
SELECT madlib.elastic_net_train( 'houses', -- Source table
'houses_en2', -- Result table
'price', -- Dependent variable
'array[tax, bath, size]', -- Independent variable
'gaussian', -- Regression family
1, -- Alpha value
30000, -- Lambda value
TRUE, -- Standardize
NULL, -- Grouping column(s)
'fista', -- Optimizer
'', -- Optimizer parameters
NULL, -- Excluded columns
10000, -- Maximum iterations
1e-6 -- Tolerance value
);
-- Turn on expanded display to make it easier to read results. \x on SELECT * FROM houses_en2;Result:
-[ RECORD 1 ]-----+--------------------------------
family | gaussian
features | {tax,bath,size}
features_selected | {tax,size}
coef_nonzero | {6.94744249834,29.7137297658}
coef_all | {6.94744249834,0,29.7137297658}
intercept | 74445.7039382
log_likelihood | -1635348585.07
standardize | t
iteration_run | 151
\x off
SELECT id, price, predict, price - predict AS residual
FROM (
SELECT
houses.*,
madlib.elastic_net_gaussian_predict(
m.coef_all, -- All coefficients
m.intercept, -- Intercept
ARRAY[tax,bath,size] -- All features
) AS predict
FROM houses, houses_en2 m) s
ORDER BY id;
\x off
SELECT id, price, predict, price - predict AS residual
FROM (
SELECT
houses.*,
madlib.elastic_net_gaussian_predict(
m.coef_nonzero, -- Non-zero coefficients
m.intercept, -- Intercept
ARRAY[tax,size] -- Features corresponding to non-zero coefficients
) AS predict
FROM houses, houses_en2 m) s
ORDER BY id;
The two queries above will result in same residuals: id | price | predict | residual ----+--------+------------------+------------------- 1 | 50000 | 101424.266931887 | -51424.2669318866 2 | 85000 | 123636.877531235 | -38636.877531235 3 | 22500 | 106081.206339915 | -83581.2063399148 4 | 90000 | 119117.827607296 | -29117.8276072958 5 | 133000 | 128186.922684709 | 4813.0773152912 6 | 90500 | 108190.009718915 | -17690.009718915 7 | 260000 | 157119.312909723 | 102880.687090277 8 | 142500 | 113935.028663057 | 28564.9713369428 9 | 160000 | 131799.592783846 | 28200.4072161544 10 | 240000 | 182913.598378673 | 57086.4016213268 11 | 87000 | 116583.600144218 | -29583.6001442184 12 | 118600 | 122842.722992761 | -4242.7229927608 13 | 140000 | 148278.940070862 | -8278.94007086201 14 | 148000 | 134883.191046754 | 13116.8089532462 15 | 65000 | 122046.449722531 | -57046.449722531 16 | 91000 | 118423.083357462 | -27423.0833574618 17 | 132300 | 127492.178434875 | 4807.8215651252 18 | 91100 | 106800.521219247 | -15700.521219247 19 | 260011 | 156424.568659889 | 103586.431340111 20 | 141800 | 114629.772912891 | 27170.2270871088 21 | 160900 | 132285.913758729 | 28614.0862412706 22 | 239000 | 182357.802978806 | 56642.197021194 23 | 81010 | 116166.753594318 | -35156.753594318 24 | 117910 | 122634.299717811 | -4724.29971781059 25 | 141100 | 148973.684320696 | -7873.68432069599 26 | 148011 | 136272.679546422 | 11738.3204535782 27 | 66000 | 122741.193972365 | -56741.193972365 (27 rows)
DROP TABLE IF EXISTS houses_en3, houses_en3_summary, houses_en3_cv;
SELECT madlib.elastic_net_train( 'houses', -- Source table
'houses_en3', -- Result table
'price', -- Dependent variable
'array[tax, bath, size]', -- Independent variable
'gaussian', -- Regression family
0.5, -- Alpha value
0.1, -- Lambda value
TRUE, -- Standardize
NULL, -- Grouping column(s)
'fista', -- Optimizer
$$ n_folds = 3, -- Cross validation parameters
validation_result=houses_en3_cv,
n_lambdas = 3,
alpha = {0, 0.1, 1}
$$,
NULL, -- Excluded columns
10000, -- Maximum iterations
1e-6 -- Tolerance value
);
SELECT * FROM houses_en3;
-[ RECORD 1 ]-----+--------------------------------------------
family | gaussian
features | {tax,bath,size}
features_selected | {tax,bath,size}
coef_nonzero | {22.4584783679,11657.0825871,52.1622899664}
coef_all | {22.4584783679,11657.0825871,52.1622899664}
intercept | -5067.27288499
log_likelihood | -543193170.15
standardize | t
iteration_run | 392
SELECT * FROM houses_en3_cv ORDER BY lambda_value DESC, alpha ASC;
alpha | lambda_value | mean | std
------+--------------+---------------------+--------------------
0 | 100000 | -1.41777698585e+110 | 1.80536123195e+110
0.1 | 100000 | -1.19953054719e+107 | 1.72846143163e+107
1 | 100000 | -4175743937.91 | 2485189261.38
0 | 100 | -4054694238.18 | 2424765457.66
0.1 | 100 | -4041768667.28 | 2418294966.72
1 | 100 | -1458791218.11 | 483327430.802
0 | 0.1 | -1442293698.38 | 426795110.876
0.1 | 0.1 | -1442705511.6 | 429680202.16
| 1 | 0.1 | -1459206061.39 | 485107796.02
(9 rows)
elastic_net_train() on a subset of the data with a limited max_iter before applying it to the full data set with a large max_iter. In the pre-run, you can adjust the parameters to get the best performance and then apply the best set of parameters to the whole data set.Elastic net regularization seeks to find a weight vector that, for any given training example set, minimizes:
![\[\min_{w \in R^N} L(w) + \lambda \left(\frac{(1-\alpha)}{2} \|w\|_2^2 + \alpha \|w\|_1 \right)\]](form_82.png)
where 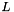 is the metric function that the user wants to minimize. Here
is the metric function that the user wants to minimize. Here ![$ \alpha \in [0,1] $](form_84.png) and
and 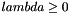 . If
. If 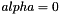 , we have the ridge regularization (known also as Tikhonov regularization), and if
, we have the ridge regularization (known also as Tikhonov regularization), and if 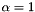 , we have the LASSO regularization.
, we have the LASSO regularization.
For the Gaussian response family (or linear model), we have
![\[L(\vec{w}) = \frac{1}{2}\left[\frac{1}{M} \sum_{m=1}^M (w^{t} x_m + w_{0} - y_m)^2 \right] \]](form_88.png)
For the Binomial response family (or logistic model), we have
![\[ L(\vec{w}) = \sum_{m=1}^M\left[y_m \log\left(1 + e^{-(w_0 + \vec{w}\cdot\vec{x}_m)}\right) + (1-y_m) \log\left(1 + e^{w_0 + \vec{w}\cdot\vec{x}_m}\right)\right]\ , \]](form_89.png)
where 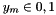 .
.
To get better convergence, one can rescale the value of each element of x
![\[ x' \leftarrow \frac{x - \bar{x}}{\sigma_x} \]](form_91.png)
and for Gaussian case we also let
![\[y' \leftarrow y - \bar{y} \]](form_92.png)
and then minimize with the regularization terms. At the end of the calculation, the orginal scales will be restored and an intercept term will be obtained at the same time as a by-product.
Note that fitting after scaling is not equivalent to directly fitting.
[1] Elastic net regularization, http://en.wikipedia.org/wiki/Elastic_net_regularization
[2] Beck, A. and M. Teboulle (2009), A fast iterative shrinkage-thresholding algorithm for linear inverse problems. SIAM J. on Imaging Sciences 2(1), 183-202.
[3] Shai Shalev-Shwartz and Ambuj Tewari, Stochastic Methods for L1 Regularized Loss Minimization. Proceedings of the 26th International Conference on Machine Learning, Montreal, Canada, 2009.
[4] Stochastic gradient descent, https://en.wikipedia.org/wiki/Stochastic_gradient_descent
File elastic_net.sql_in documenting the SQL functions.